String manipulations are an essential part of Data Science. The latest release of Vaex adds incredibly fast and memory efficient support for all common string manipulations. Compared to Pandas, the most popular DataFrame library in the Python ecosystem, string operations are up to ~30–100x faster on your quadcore laptop, and up to a 1000 times faster on a 32 core machine.
While Pandas is largely responsible for the popularity of Python in data science, it is eager for memory. As data gets bigger, you will have to be careful not to have a MemoryError ruin your day. Switching to a more powerful machine may solve some memory issues, but now your 31 cores sit idle. Pandas will only use one out of the 32 cores of your fancy machine. With Vaex, all string operations are out of core, executed in parallel, and lazily evaluated, allowing you to crunch through a billion-row dataset effortlessly.
“Almost 1000x faster string processing, corresponding to 1 minute versus 15 hours! ”
Vaex and superstrings
Vaex is now moving beyond fast number crunching, expanding into the (super) string realm. The latest release includes almost all Pandas’ string operations. You can now do all your string operations lazily, as demonstrated by counting the string lengths in a DataFrame containing one googol (1 followed by 100 zeros) rows.
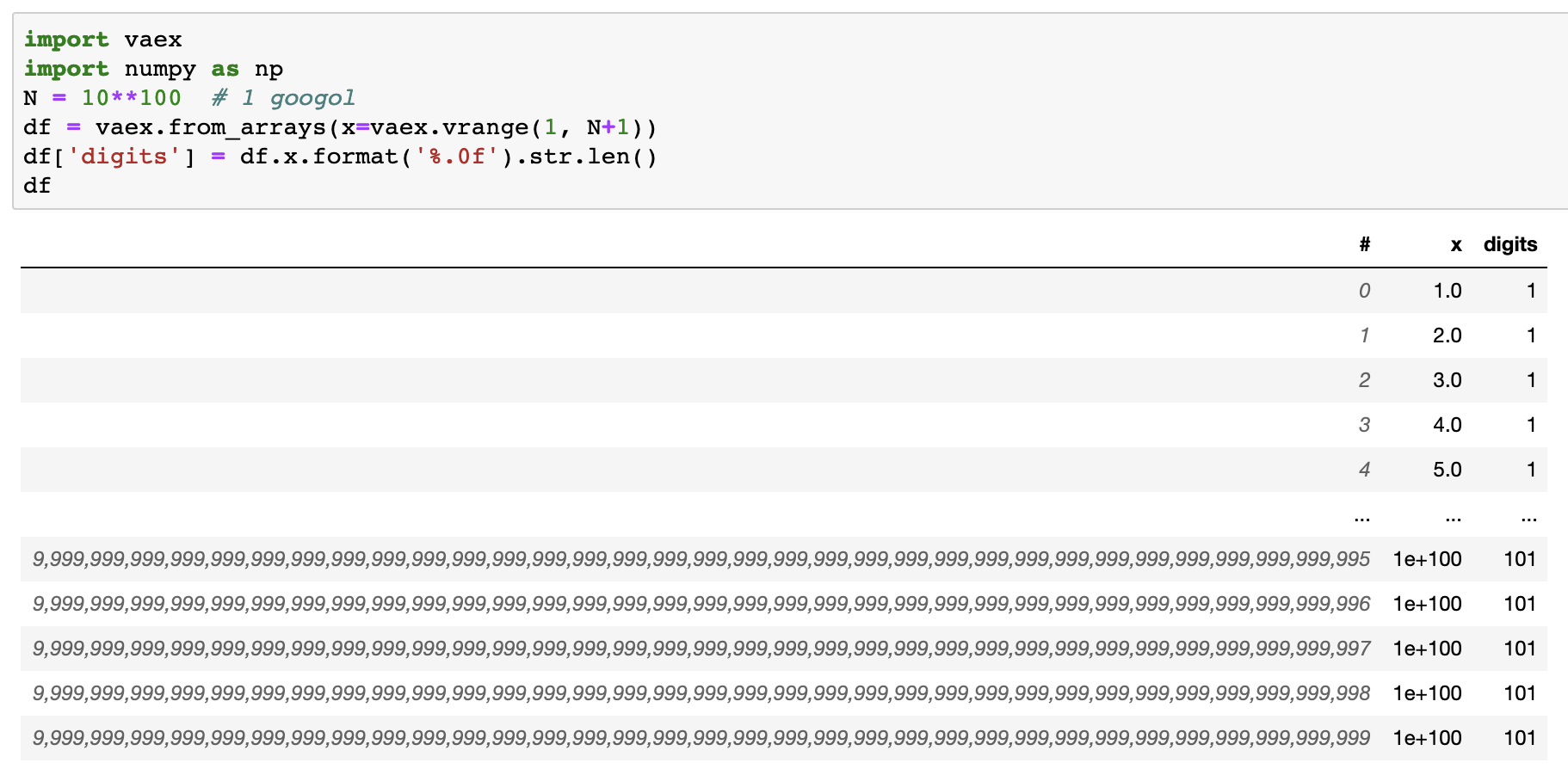
A googol length DataFrame demonstrates the advantage of lazy evaluations. When experimenting with regular expressions on your badly malformed input data, you don’t have to wait for a minute to see if all your operations went fine. Instead, you apply your operations, print out the dataframe, and directly see a preview (head and tail), without any delay. Only the values you see are evaluated.
Performance benchmarks
Eventually, you have to process all the data. The lazy evaluation does not accelerate the operations, it only postpones. If you need to export the data or pass it to a machine learning library, it needs to happen eventually right? So how well does vaex perform? Let us take an artificial dataset with 100 million rows (see the appendix for details), and perform the same operations with Pandas and Vaex (Spark comes later):
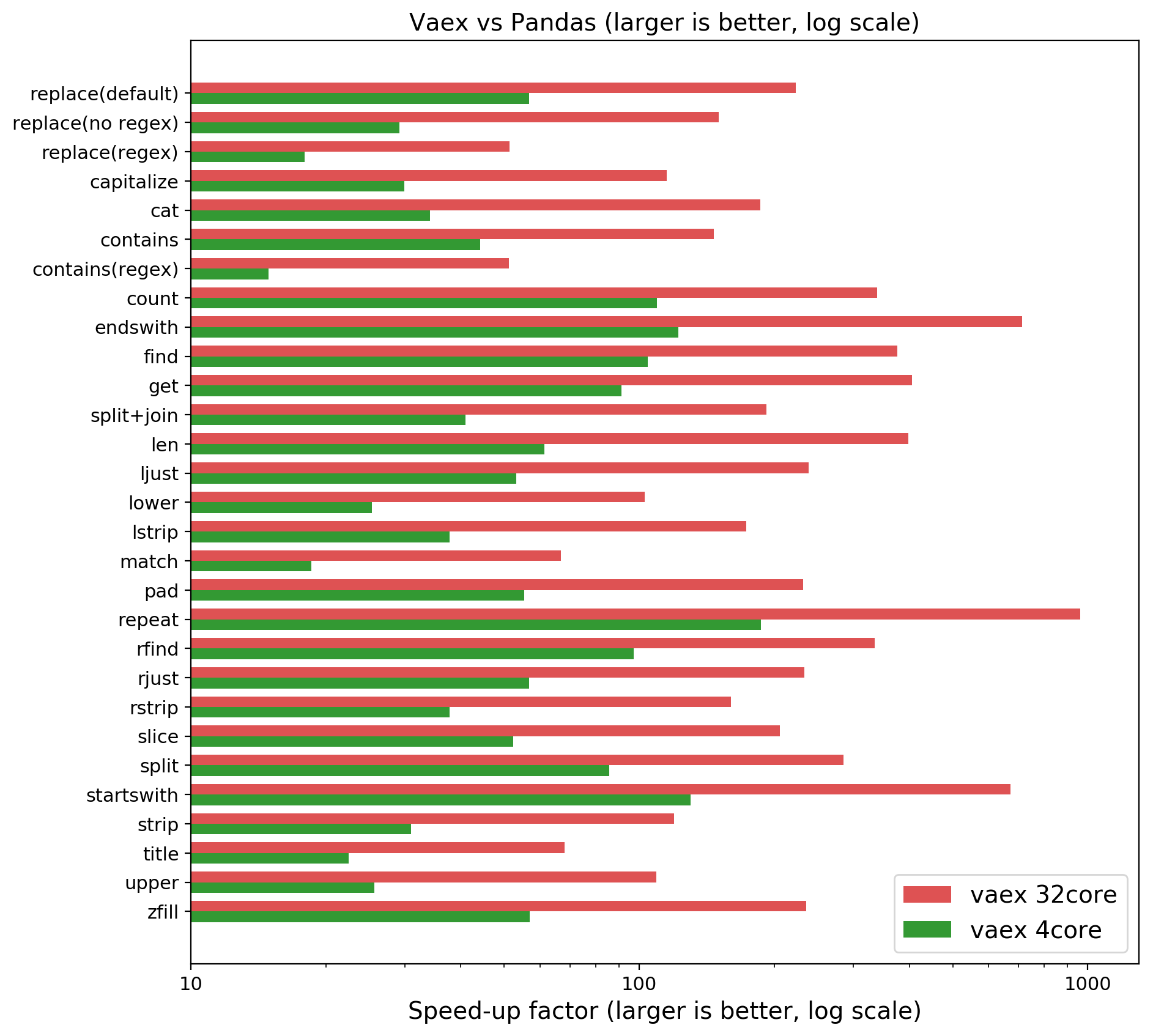 NOTE: Larger is better, x is logarithmic. Speedup of Vaex compared to Pandas on a quadcore laptop or 32 core machine. On your laptop you may expect 30–100x, on an AWS h1.8xlarge (32 core) we measured close to a 1000x speedup!
NOTE: Larger is better, x is logarithmic. Speedup of Vaex compared to Pandas on a quadcore laptop or 32 core machine. On your laptop you may expect 30–100x, on an AWS h1.8xlarge (32 core) we measured close to a 1000x speedup!
Vaex performs up to ~190x faster on my quadcore laptop, and even up to ~1000x faster on an AWS h1.x8large machine! The slowest operations are the regular expressions, which is expected. Regular expressions are CPU intensive, meaning that a large fraction of the time is spent in the operations instead of all the bookkeeping around them. In short, the overhead of Python is low here, and Python’s regular expression engine is fast already, so we ‘only’ get a factor of 10–60x in speedup.
How: The GIL, C++, and ApacheArrow
How is that even possible? Three ingredients are involved: C++, Apache Arrow and the Global Interpreter Lock GIL (GIL). In Python multithreading is hampered by the GIL, making all pure Python instructions effectively single threaded. When moving to C++, the GIL can be released and all cores of your machine will be used. To exploit this advantage, Vaex does all string operations in C++.
The next ingredient is to define an in-memory and on-disk data structure that is efficient in memory usage, and this is where Apache Arrow comes into play. Apache Arrow defines a well thought out StringArray that can be stored on disk, is friendly to the CPU, and even supports masked/null values. On top of that, it means that all projects supporting the Apache Arrow format will be able to use the same data structure without any memory copying.
What about dask?
People often ask how Dask compares to Vaex. Dask is a fantastic library that allows parallel computations for Python. We actually would love to build Vaex on top of dask in the future, but they cannot be compared. However, Vaex can be compared against dask.dataframe, a library that parallelizes Pandas using Dask. For the benchmarks we ran, dask.dataframe was actually slower than pure Pandas (~2x). Since the Pandas string operations do not release the GIL, Dask cannot effectively use multithreading, as it would for computations using numpy, which does release the GIL. A way around the GIL is to use processes in Dask. This, however, slowed down the operations by 40x compared to Pandas, which is 1300x slower (!) compared to Vaex. Most of the time is spent pickling and unpickling the strings.
While Dask is a fantastic library, dask.dataframe cannot perform magic in the strings realm. It inherits some of Pandas’ issues, and using processes can give a large overhead due to pickling. Although there might be ways to speed this up, the out of the box performance is not great for strings.
What about Spark?
Apache Spark is the library in the JVM/Java ecosystem to handle large datasets for data science. If Pandas cannot handle a particular dataset, people often resort to PySpark, the Python wrapper/binding to Spark. This is an extra hurdle if your job is to produce results, not to set up Spark locally or even in a cluster. Therefore we also benchmarked Spark against the same operations:
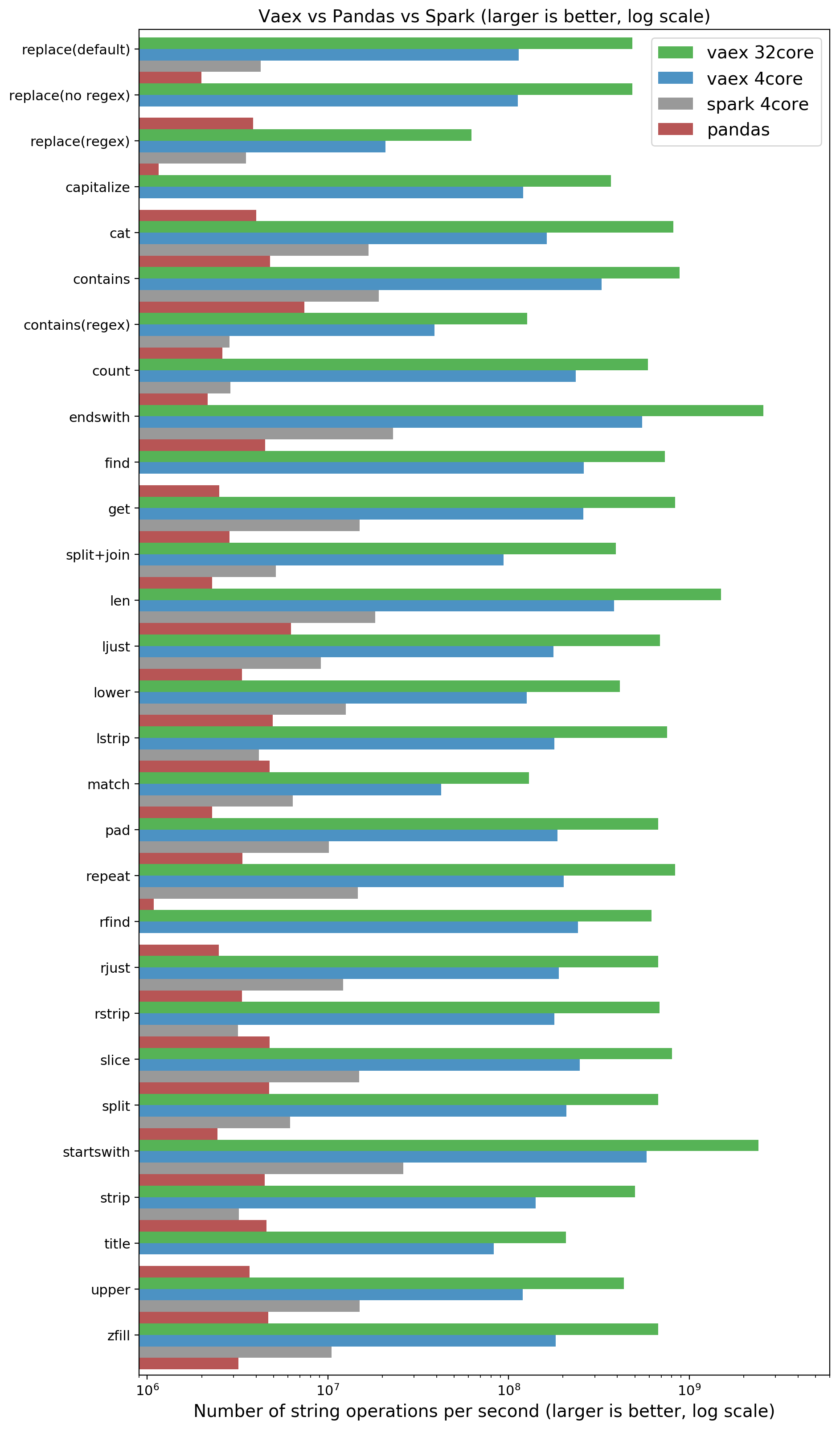 NOTE: Larger is better, x is logarithmic. Comparing Vaex, Pandas and Spark.
NOTE: Larger is better, x is logarithmic. Comparing Vaex, Pandas and Spark.
Spark performs better than Pandas, which is expected due to multithreading. We were surprised to see vaex doing so much better than Spark. Overall we can say that if you want to do interactive work on your laptop:
-
Pandas will do in the order of millions of strings per seconds (and does not scale)
-
Spark will do in the order of 10 millions of strings per second (and will scale up with the number of cores and number of machines).
-
Vaex can do in the order of 100 millions of strings per second, and will scale up with the number of cores. On a 32 core machine, we get in the order of a billion of strings per second.
Note: some operations will scale with the string length, so the absolute number may be different depending on the problem.
The future
Pandas will be around forever, its flexibility is unparalleled, and for a big part responsible for the Python’s popularity in data science. However, the Python community should have good answers when datasets become too large to handle. Dask.dataframe tries to attack large datasets by building on top of Pandas, but inherits its issues. Alternatively, nVidia’s cuDF (part of RAPIDS) attacks the performance issues by using GPU’s, but requires a modern nVidia graphics card, with even more memory constraints.
Vaex not only tries to scale up by using more CPU’s/cores and efficient C++ code, but it also takes a different approach with its expression system / lazy evaluation. Calculations and operations are done only when needed and performed in chunks, so no memory is wasted. More interestingly, the expressions that led to your results are stored. This is the foundation of vaex-ml, a new approach to doing machine learning, where pipelines become an artifact of your exploration. Stay tuned.
Conclusions
Vaex uses ApacheArrow data structures and C++ to speed up string operations by a factor of about ~30–100x on a quadcore laptop, and up to 1000x on a 32 core machine. Nearly all of Pandas’ string operations are supported, and memory usage is practically zero because the lazy computations are done in chunks.

-
A clap for this article or a ⭐on GitHub is appreciated.
-
Vaex has the same API as Pandas.
-
See the tutorial for the usage.
-
Submit issues if you found a missing feature or bug.
-
pip install vaex / conda install -c conda-forge vaex or read the docs
Appendix
Benchmarks are never fair, can sometimes be artificial, and are not identical to the real world performance. Everybody knows or should know this. A benchmark gives you an impression of what to expect, but especially with operations like regular expressions, it becomes tricky to compare. A benchmark is better than no benchmark. In any case, if you want to reproduce these results, you can do so by using this script. The Spark benchmarks can be found at this location.
